P4SPR – Detection of hemagglutinin
Detection of hemagglutinin using Affinité’s P4SPRTM as a potential platform to monitor production of influenza vaccine
IntroductionThe influenza vaccine is manufactured each year to protect people against serious illness that require hospitalization and reduce the number of mortalities associated with influenza. Therefore, the timely production and validation of influenza vaccine lots are of utmost importance. Each vaccine lot must be evaluated for potency, which is based on the amount of the hemagglutinin (HA) content found on influenza viruses in the vaccine. Using HA, these viruses are known to infect host cells by binding to sialic acid at the end of complex glycans [1].
The current gold standard method to evaluate vaccine lots is single-radial immunodiffusion (SRID) assay, which lets viral antigens diffuse in an agarose gel containing antibodies specific to the antigens applied. Then, the size of the precipitated zone is measured, which is correlated to the amount of antigens in the sample by using a set of calibrated reference antigens [2]. Although SRID has been used since 1978, it has a few disadvantages. First, it is an off-line method, as samples must be removed from the production line to be analysed through a separate process. Secondly, reference sera and antigen reagents required for SRID may take 6 months to prepare and cause delays in releasing vaccine lots to the public [3]. Furthermore, SRID also has low analytical accuracy and throughput capacity [4], and it takes about 22 h to perform the SRID [5]. Therefore, there is an urgent need to develop a more rapid method that can evaluate influenza and other vaccine lot potencies in anticipation for flu seasons or any high viral activities, and especially during a pandemic outbreak.
Although there are other immunoassay and chromatography-based methods that can determine the HA content in vaccine preparations, they lack the ability to perform in-process analysis of the virus particle production level. Immunoassays are also known to be time-consuming and labourintensive. Surface plasmon resonance (SPR), on the other hand, produces real-time data within minutes to hours and is label-free. Upon optimization of surface chemistry and careful selection of the capture biomolecule, detection of the target biomolecule can be achieved with high sensitivity and reproducibility. Moreover, Affinité Instruments’ selection of portable and compact SPR instruments would render the on-line, real-time monitoring of cell-based viral particle production for manufacturing vaccines a reality.
In this application note, the generation of a calibration curve of HA samples will be used as a proof-of-concept experiment that Affinité’s P4SPRTM can potentially be used as a monitoring device for influenza vaccine production.
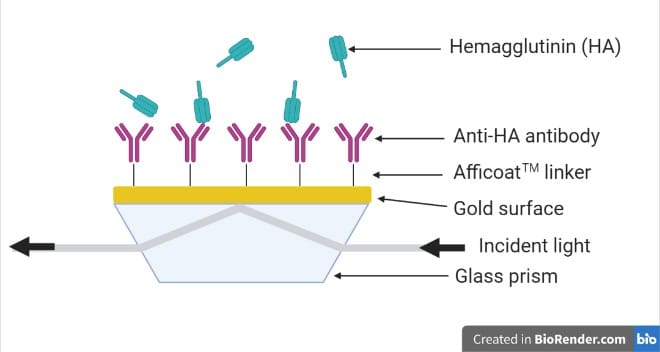
Experimental proceduresSystem setup and sensor chip functionalization. A quad inlet P4SPR™ equipped with 4 separate microfluidic channels was used for this study (Figure 1).
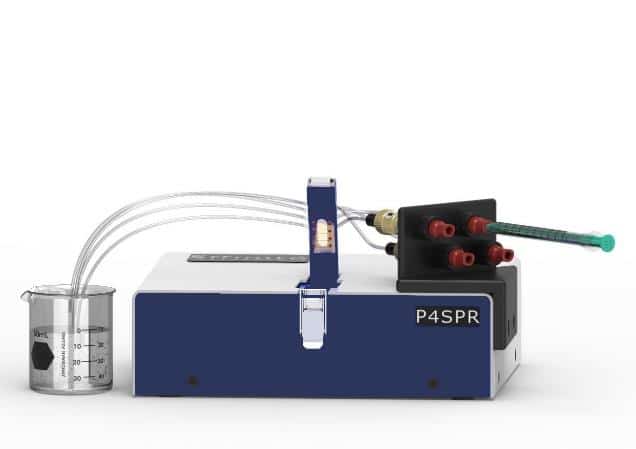

Initially, distilled water was run through the SPR device for 60s to ensure stability of baseline. Then, to reduce non-specific binding, a gold sensor surface modified with our proprietary AfficoatTM was used, and all the following protein samples and reagents were manually injected into the P4SPR (see Figure 1). To activate the sensor surface for capture antibody immobilization, it was treated with EDC/NHS and subsequently washed with sodium acetate. The capture antibody for HA (10 µg/mL) was immobilized onto the surface for twenty minutes. Another wash using sodium acetate was then carried out. To reduce non-specific binding, the active sites on the sensor surface were blocked by injecting 1 M ethanolamine, pH 8.5, for ten minutes. The sensor surface was then washed with running buffer. Figure 2 depicts the experimental setup on the gold sensor chip.
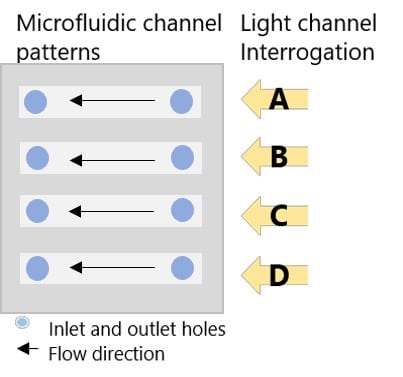
Standard curve of HA. Subsequently, HA, prepared in running buffer, was titrated in channels A to C from 0.5 to 25 µg/mL using 10 minutes for each injection. Figure 3 shows the microfluidic channel pattern that is placed on top of the sensor chip. As a control, channel D was dedicated to injections of running buffer. The SPR shifts for each set of HA concentrations and running buffer were saved in the P4SPR control software.
 Results and Discussion
Results and DiscussionThe initial P4SPRTM washing, stabilization, and sensor chip functionalization steps required about 38 min. The injections of HA samples to create the standard curve required about 1 h. Therefore, the total time it took to generate the standard curve was about 1 h and 40 min.
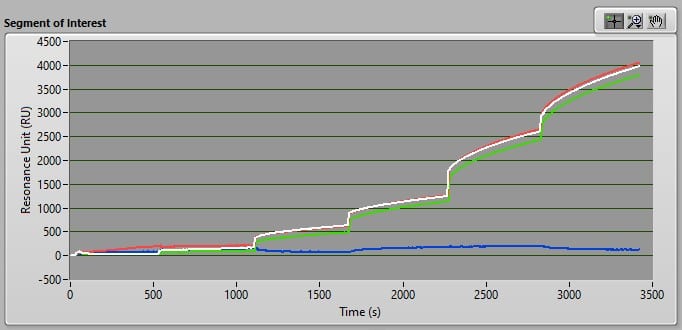
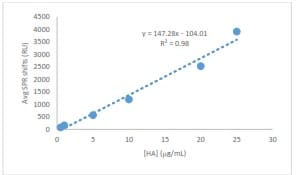
Binding of HA to immobilized anti-HA antibody was demonstrated by a proportional increase in resonance signal with each injection of HA concentration (Figure 4). At each HA concentration, the triplicate resonance signals were averaged and plotted in Excel. The resulting standard curve generated an R2 of 0.98, indicating a high degree of linearity between concentrations of 0.5-25 µg/mL (Figure 5).
Conclusions
This study demonstrates that the P4SPRTM can be used to detect the amount of HA in a sample through immobilized anti-HA antibodies onto our proprietary AfficoatTM sensor chips and by using a standard curve. The sensor surface is ready to be used within 40 min and each sample requires only 10 minutes to obtain data. Through further assay optimization, the P4SPR will be able to evaluate the viral potency in real production and purified vaccine samples.
The Affinité AdvantageAffinité instruments’ P4SPRTM is lightweight, compact, and portable. The minimal requirements for setup space and signal stabilization are the P4SPR’s key features. During the vaccine production process, it would be advantageous to install an online SPR instrument to evaluate vaccine lots in real-time.
References1. Emi Suenaga, Hiroshi Mizuno & Penmetcha K.R. Kumar. Influenza virus surveillance using surface plasmon resonance. Virulence, 2012, 3, 464-470.
2. M.S. Williams. Single-radialimmunodiffusion as an in vitro potency assay for human inactivated viral vaccines. Vet. Microbiol., 1993, 37, 253-262.
3. Laurent Durous, Thomas Julien, Blandine Padey, Aurélien Traversier, Manuel RosaCalatrava, Loïc J. Blum, Christophe A. Marquette, Emma Petiot. SPRi-based hemagglutinin quantitative assay for influenza vaccine production monitoring. Vaccine, 2019, 37, 1614-1621.
4. Carl-Fredrik Mandenius, Ronghui Wang, Anna Alden Gunnar Bergstrom, Sabine Thebault, Charles Lutsch, Sten Ohlson. Monitoring of influenza virus hemagglutinin in process samples using weak affinity ligands and surface plasmon resonance. Anal. Chim. Acta, 2008, 623, 66- 75.
5. C. Estmer Nilsson, S. Abbas, M. Bennemo, A. Larsson, M.D. Hämäläinen, Å. FrostellKarlsson. A novel assay for influenza virus quantification using surface plasmon resonance. Vaccine, 2010, 28, 759–766.
AcknowledgementsWe would like to deeply thank Cristina Silva from Dr. Amine Kamen’s lab at McGill University for providing the samples. We would also like to thank Abdelhadi Djaileb from Dr. Jean-François Masson’s lab at Université de Montréal for the collection of these data. Figure 1 was produced using Biorender.com. About Affinité Instruments Established in 2015 as a spin-off of the Université de Montréal, Affinité instruments’ foundation is built on deep knowledge accrued over a decade of research in SPR. The commercialization of promising innovations is spearheaded by a diverse leadership experienced in business, science, and engineering.
About the AuthorsCory Campbell is a Ph.D. candidate in biochemistry at Concordia University and has experience using multiple SPR platforms to investigate proteinprotein interactions. Dr. April Wong is an analytical chemist and has research experience in the biosensor area for over a decade. She obtained her Ph.D. at the University of Toronto.
