Beckman Coulter – Liposome Characterisation Application Note
Techniques of the Coulter Principal and Dynamic Light Scattering
ABSTRACT
The following application note is a valuable resource for any researcher working with particle characterisation. Sub-micron liposomes are characterised by Dynamic Light Scattering (DLS) and the Coulter Principle, using the latest technology and instruments from Beckman Coulter. Both quality control scientists and exploratory research scientists will benefit from learning how the or thogonal techniques of DLS and the Coulter Principle can accurately validate particle size, while eliminating spurious results.
INTRODUCTION
Liposomes are special lipid vesicles ranging from 50 nm to 5 µm in diameter. Liposomes have a wide range of applications in numerous industries including textiles1, cosmetics industry2, the medical industry3, and biological research4,5. The encapsulation of genes and molecules inside liposomes offers an efficient drug delivery mechanism6,7. In biological research, the reconstitution and study of membrane proteins in liposomes offer a simplified cellular system allowing for detailed biophysical studies. When forming liposomes, determining the final count of total liposomes is useful for successful and repeatable results.
Determining the exact liposome concentration from both a mass perspective (grams/litre) and a count perspective (particles/litre) can aid in precision dosing of therapeutic compounds in the medical industries. For biomimetic systems, determining the total number of liposomes can be crucial for efficient and functional reconstitution of membrane proteins into liposomes.
Sizing and characterising in the submicron region can generate particle population statistics through either an ensemble or a discrete approach. Ensemble techniques include sedimentation, sieving, and light scattering (dynamic and static). Discrete characterising techniques include electron microscopy, optical microscopy, and the Coulter Principle. Sedimentation and sieving techniques have very limited accuracy and require long periods of time for each measurement. DLS rapidly delivers average diameter and polydispersity of the sample, however, as an ensemble technique, concentration information is fundamentally unavailable. Discrete techniques, like electron and optical microscopy, give accurate size and shape for particles. Only a small portion of the particles is analysed, while microscopy measurements are tedious, subject to user bias, and still lacking concentration information. Using the Coulter Principle with DLS provides additional details including concentration, while verifying the size information derived from DLS on a rigorous, particle-by-particle basis.
METHOD
DelsaMax CORE Analysis
Liposomes were purchased for analysis by Librede, a leading company in lipid development. The liposomes were analysed within three weeks of production. The liposomes were made of lipid type DPhPC (1,2-diphytanoyl-sn-glycero-3-phosphocholine); the suspension buffer comprised 150 mM NaCl, 5 mM CaCl2, 10 mM HEPES, and pH 8.0. The initial concentration was 0.8 mg/mL. For DLS analysis on the DelsaMax CORE, the liposomes were diluted 1:100 in deionised, sub-micron filtered water—giving a final concentration of 0.008 mg/mL, and analysed in the disposable cyclic polyolefin cuvette using 20 µl of sample. The liposome sample was run five times; each run consisted of three acquisitions that lasted five seconds/acquisition. The set cell temperature was 25° C.
All sample runs had a Sum-of-Squares (SOS) fit of less than 10, indicating a clean sample without any contamination from dust or bubbles. Diameter and polydispersity reported are from cumulant analysis, which assumes a monomodal sample; regularisation analysis (representative data shown in Figure 2) indicates that the sample is monomodal.
DelsaMax PRO Analysis
The DPhPC liposomes from Librede were also analyzed on the DelsaMax PRO, which offers simultaneous size and zeta potential. As a comparison, generic liposomes from Lypo-Spheric™ were also diluted in water and run. The DPhPC liposomes were run in the same conditions as above (0.008 mg/mL), while the generic liposomes were run at ~0.05 mg/mL in water. The samples were injected into the flow cells (~200 µl/sample); each sample was analysed a total of five times. The zeta potential collection period was 20 seconds, during which DLS data was collected as well (10 runs, two seconds/run). All samples were run at 25° C. The representative size and zeta potential are shown in Figure 3 while the quantitative data is shown in Table 2.
Multisizer 4e COULTER COUNTER Analysis
The same stock liposome solution used in the DelsaMax CORE analysis was used for the Multisizer 4e analysis. The liposomes were diluted with ISOTON II Diluent (Beckman Coulter, Inc.) and run on a Multisizer 4e instrument equipped with 10 micron aperture; 5 µl of stock liposome solution (0.8 mg/mL) was diluted with 19.995 mL of ISOTON II (1:4,000 dilution). The 10 µm aperture was installed for the analysis and the analytical range was 0.2 to 6 µm with 400 size bins logarithmically spaced. The analysis was run in volumetric mode in order to determine liposome molarity; 10 µl of sample were analysed in total. Ten separate runs were conducted.
DISCUSSION
The study of membrane proteins in artificial systems provides unique insight into the function of these important proteins. During reconstitution of proteins into liposomes, the protein to lipid ratio is an important metric to prevent protein aggregation. During reconstitution, pre-formed liposomes are often softened with mild detergents. These detergent-saturated liposomes are then mixed with a detergentsolubilised purified-membrane protein. Knowing the exact number of liposomes can help control the reconstitution efficiency and control the protein to lipid ratio, allowing the researcher a greater control over the biological environment.
The Multisizer 4e COULTER COUNTER (MS4e) has the largest dynamic range of any instrument based on the Coulter Principle, spanning from 200 nm up to 1,600 microns. Biologically relevant samples are typically in salt solutions that are perfect for running on the MS4e. By analysing the entire sample particle-by-particle, any outlier can be easily detected and quantified; outlier detection is imperative in pharmaceutical formulation because one large aggregated outlier could cause blood vessel blockage and embolism. In Figure 1, particle size is plotted in bins with width less than 10 nm in size. There are 400 total bins across the diameter range (x-axis). However, because the MS4e digitally acquires and saves the pulse height and width data, reanalysis can be done with any width of analysis. This allows the user to zoom in on an area of interest and have particle bin sizes less than 0.1 nm in diameter, as seen in Figure 4.
The DelsaMax CORE uses a complementary, light-scattering size detection or thogonal technique to determine the average diameter of the sample. The analysis on the DelsaMax CORE takes as little as one second and can easily verify the sizing data from the MS4e, as shown in Figure 2. The average diameter of the liposomes in the DelsaMax CORE is reported at 320.2 ± 19.8 nm—very close to the mean diameter of the liposomes from the MS4e (343 nm) and much larger than the median size (292 nm) from the MS4e (Table 1). Large particles in the liposome mixture scatter exponentially more light, skewing the average size distribution data to higher values.
Based on 10 runs, the final liposome particle count in the diluted sample run in the MS4e was 1.5132 ± 0.144 * 1010 liposomes/mL. The final liposome particle concentration can be easily visualised in the MS4e using the cumulative plot function (data not shown). Given a dilution of 1:4,000, the stock liposome concentration had 6.0528 ± 0.574 * 1013 liposomes/mL (100.5 femtomolar).
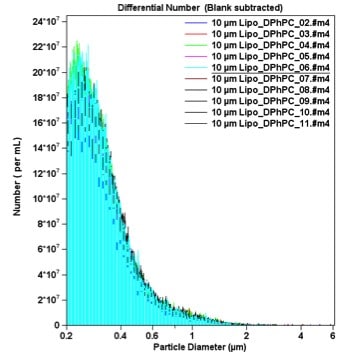
Figure 1. Differential Number Count of Liposomes. Ten runs were conducted and a blank was used for background subtraction. A 10-micron aperture was used in order to determine the size down to 200 nm. The majority of liposomes were between 200 to 400 nm; several larger species greater than one micron in size skew up the mean liposome diameter. The y-axis is given in terms of particle concentration.
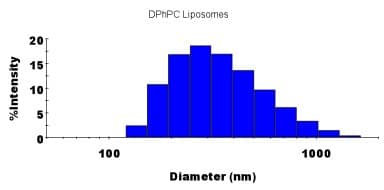
Figure 2. Dynamic Light Scattering Measurement of the Average Diameter of the DPhPC Liposomes. Regularisation analysis was used to fit the auto-correlation function; if the sample was multimodal in size, the regularisation plot above would contain multiple peaks.
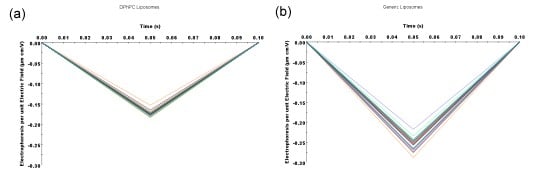
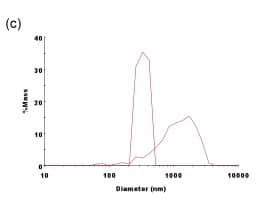
Figure 3. Comparison of Generic Liposomes vs. DPhPC-Extruded Liposomes on the DelsaMax PRO. (a) The raw mobility data from all 31 zeta potential detectors for the DPhPC liposomes; and (b) the raw mobility data from all 31 zeta potential detectors for the generic liposomes. Thirty-one independent zeta potential measurements verify the accuracy of the measurement while greatly decreasing the time it takes for each measurement. (c) Diameter comparison of the two types of liposomes. Note that larger particles are clearly present in the generic liposomes.
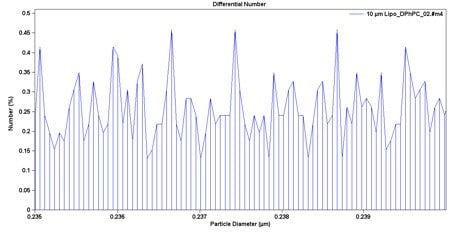
Figure 4. A Zoomed-In View of the MS4e Data. Because the pulse data is saved digitally, particle size bin width can be recalculated to have any width that is needed, with minimum-width size smaller than 0.1 nm/bin. In this plot, the percent of liposome particles from 235 nm to 240 nm is displayed.
Table 1. The two reported mean diameters are within 10%, supporting the veracity of each particle characterisation method.
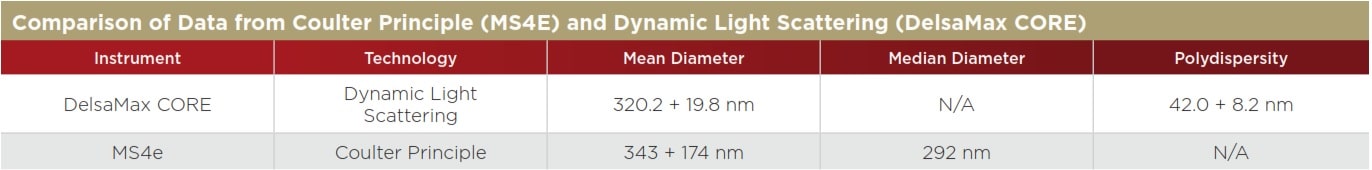
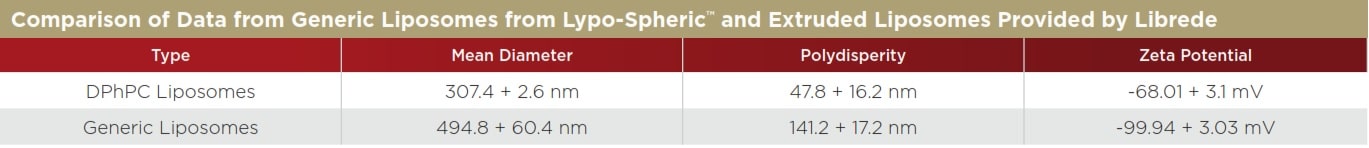
Table 2. Note that the extruded liposomes from Librede have a much smaller polydispersity, indicating that the sample is much more uniform in size. The zeta potential of both liposomes is highly negative, indicating that both are quite stable in water. Most particles with a zeta potential more negative than -30 mV are stable based on electrostatic repulsion.
REFERENCES
1. Nelson G. Application of microencapsulation in textiles. International Journal of Pharmaceutics. 242.1; 55–62: (2002).
2. Memoli A et al. Liposomes in cosmetics: Which kind of phospholipid? Which loading method? Journal–Society of Cosmetic Chemists. 44; 123: (1993).
3. Lasic D D and Papahadjopoulos D, eds. Medical applications of liposomes. (Elsevier Science, 1998).
4. Kumar K P S, Bhowmik D, and Deb L. Recent Trends in Liposomes Used As Novel Drug Delivery System. The Pharma Innovation. 1; 29–38: (2012).
5. Jølck R I et al. Engineering liposomes and nanoparticles for biological targeting. Biofunctionalization of Polymers and their Applications. Springer Berlin Heidelberg. 251–280: (2011).
6. Kaneda Y. Virosomes: evolution of the liposome as a targeted drug delivery system. Advanced drug delivery reviews. 43.2; 197–205: (2000).
7. Cao Z et al. Reversible Cell-Specific Drug Delivery with Aptamer-Functionalized Liposomes. Angewandte Chemie International Edition. 48.35; 6494–6498: (2009).
